Catalysts: Chemistry’s Facilitators
In organic and inorganic instances alike, a chemical reaction is defined as a process of breaking and/or making bonds that has two distinct sides, the reactant side and the product side. A catalyst is a substance that increases the rate of a reaction without being chemically changed. That is, a catalyst is not consumed. For instance, in the Haber process, an iron catalyst is used to make ammonia from hydrogen and nitrogen and the iron is not used up. These inorganic catalysts are used to improve the yield of a product many-fold over. We also find catalysts in nature, called enzymes. Enzymes are catalysts made up of proteins. In a reaction, all reactants must overcome a barrier of energy, called the activation energy, to form products.
1
Enzymes are the biological molecules that lower this activation energy necessary for chemical reactions to begin and, in turn, increase the rate of these reactions. Figure 1 shows a reaction energy diagram with and without a catalyst and illustrates this phenomenon.
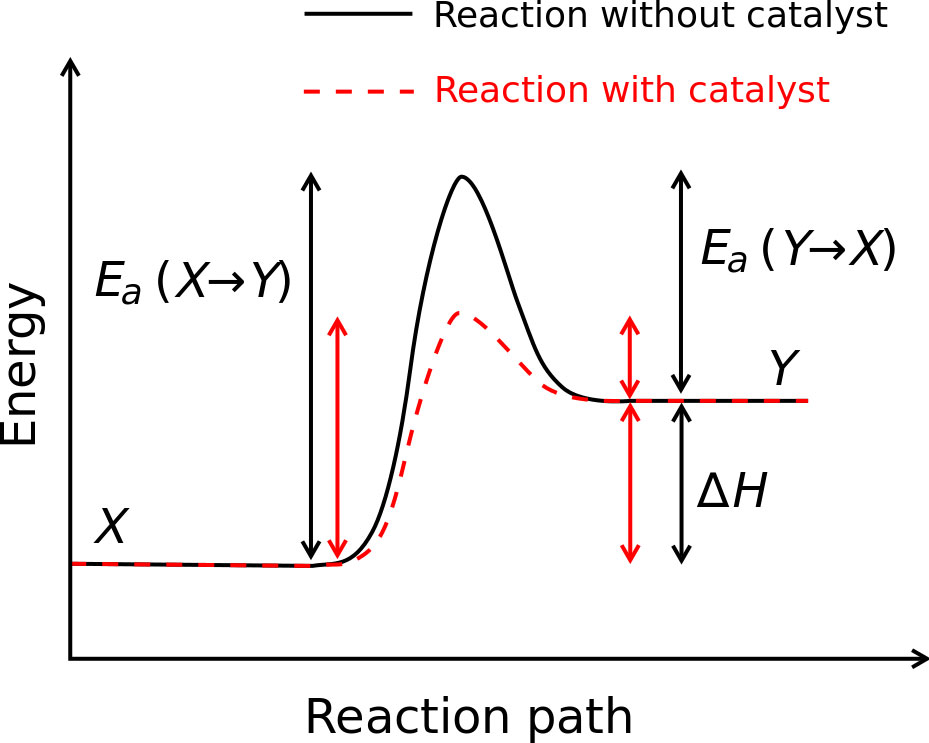
Figure 1: https://en.wikipedia.org/wiki/File:Activation_energy.svg
An Introduction to Enzymes
Though their activity had been studied during the 17
th
and 18
th
centuries, enzymes and their biochemical nature were relatively unknown up until the early 1900s. Some scientists believed proteins were simply carriers for the true enzymes, which performed catalysis. In fact, it was not until around 1930, that pure proteins were shown to be capable of catalysis through the work John Northrop and Wendell Stanley conducted on pepsin, trypsin and chymotrypsin.
2
In the years since, enzymology, the study of enzymes—their structure, function and kinetics—has become a cornerstone of biochemistry. This subject is particularly important because enzymes serve as conduits for the accelerated reaction times needed to support life.
3
In particular, enzymes play an integral role in cellular function, catalyzing more than 5,000 known reaction types.
4
Let’s explore one example. Proteases are a particular class of enzymes that cleave peptide bonds—the bonds that hold amino acids together. That is, proteases cut proteins. When we eat food our body uses a number of proteases to break down the proteins we ingest. One such example is trypsin, a protease that is secreted in the part of the stomach called the duodenum. Trypsin cuts proteins after particular amino acids, lysine and arginine. These peptide products are then broken down by other proteases until they are able to be absorbed into the blood stream. This example shows us that enzymes are not only biologically useful, but have specificity—only acting after certain amino acids—a topic we will explore more in the coming sections.
Moreover, this example allows us to gain insight into how enzymes provide a pathway for reactions to occur. To build a protein it takes energy. Amino acids must be connected one by one until the correct sequence is assembled. The final product—the protein itself—is therefore at a higher energy state than all the separate and individual amino acids it took to build it. And yet, these high-energy proteins do not spontaneously degrade into individual amino acids. To do that, they need a pathway to get to a lower energy level. Proteases provide that pathway and catalyze protein cleavage.
Enzymes operate essentially the same way inorganic catalysts operate. They both accelerate reaction rates by lowering the activation energy. For the Haber process, an inorganic process, and for protease cleavage, an enzyme-mediated process, the activation energies are lowered and the reaction rates are increased. For both the inorganic catalyst and the enzyme, the net effect is the same—dramatic increases in the rate of conversion of reactant to product, sometimes making the rate millions of times faster.
Yet, unlike inorganic catalysts, enzymes function in biological environments and have a number of special characteristics. In particular, enzymes allow for greater reaction specificity, greater capacity for regulation, and milder reaction conditions. Enzymes can also couple an energy releasing catalysis, such as the hydrolysis of ATP, with an energy consuming catalysis to move an unfavorable reaction forward. In this case, coupling a reaction allows enzymes to create products of a higher energy than the reactants. Each of these characteristics contributes to making enzymes unique and adaptable structures.
Enzyme Structure
Proteins are polymers of amino acids. To be biologically relevant, a string of amino acids forms a globular protein through numerous interactions. Although they range in size from tens to thousands of residues, proteins are usually between 1-100 nanometers large.
5
Protein structure is important because it creates enzyme specificity. A string of amino acids ends up forming highly defined enzymes with specific functions.
We will start with an example of a chemical polymer with no defined structure to understand how specificity arises. Chemical polymers have no sequence variation—the same monomer is added over and over. Low-density polyethylene (LDPE) is an example of a chemical polymer that we all encounter. Made from the monomer ethylene, flexible and chemically inert, LDPE is used in plastic bags, toys, flexible pipes, squeeze bottles and numerous other common household items. At its surface LDPE interacts the same way with everything it touches because its surface is identical at every point; there is no specificity inherent in its structure. The only way we impart function on LDPE is when we alter its global shape—not surface—and artificially mold it.
Biological polymers, such as proteins, are very different. They are inherently defined and specific structures. From DNA we get a defined sequence of amino acids rather than a string of the exact same monomer. This defined sequence then makes defined folds and creates defined surfaces. Unlike LDPE, the surface of a protein is not identical everywhere. Rather, it is highly varied. Part of this surface will be the binding pocket where the reactant will fit. This is how enzymes are selective: discrete sequences create defined folds, which interact with specific molecules. Now that we have reviewed a framework for understanding the protein structure and specificity, we will engage structural details.
There are four separate levels of protein structure termed primary structure, secondary structure, tertiary structure, and quaternary structure. Primary structure refers to the amino acid sequence of the polypeptide chain. The primary structure is created during translation in which a specific sequence of mRNA nucleotides is read by the ribosome, which in turn, produces a protein. It is noteworthy that the primary structure is determined by the gene that coded the protein. That is, the protein’s DNA is transcribed into RNA, which is then translated into protein.
Secondary structure refers to highly organized local structures. There are two main types of secondary structure, the alpha helix and the beta sheet, which Linus Pauling, a two-time Nobel Prize winner, proposed in 1951.
6
Alpha helices and beta sheets both have a regular pattern of hydrogen bonds between peptide chains. The alpha helix is a righthand-coiled structure while the beta sheet is a twisted, pleated sheet with either parallel or antiparallel stands of polypeptide hydrogen bonded together.
Tertiary structure refers to the interactions between the secondary structures, which give rise to the three dimensional structure of the protein. This folding is driven by hydrophobic interactions in which fatty amino acids collapse away from the surface of the protein—like how oil beads in water. In addition to this, tertiary structure is characterized by the presence of salt bridges, hydrogen bonds, and packing of disulfide bonds. Lastly, quaternary structure refers to the overall three-dimensional structure of a multi-subunit protein. Quaternary structures are stabilized by non-covalent interactions and disulfide bonds, which link subunits together.
These levels of protein structure will be generalized again for understanding. The amino acids that make up proteins are first chained together in a defined sequence that is determined by DNA. This defined sequence then has all the information necessary to make defined folds and defined binding pockets—the places where reactants settle for catalysis. These defined folds have limitless possible arrangements and limitless possible functions. This is how enzymes can do everything from helping digest food—remember trypsin—to packaging DNA. Enzymes carry out all of the functions necessary for life.
Enzyme Catalysis
Before a reaction can be catalyzed, enzymes have to bind their substrates. Several models to explain how this actually occurs have been proposed. Initially, Emil Fischer’s 1894 lock and key model was widely accepted. It held that the enzyme and the substrate were like puzzle pieces with complementary shapes that fit together perfectly. The model explained how enzymes could distinguish substrates from other molecules, but overlooked how the stability of the transition state is achieved.
7
In 1958, Daniel Koshland proposed a new model known as the induced fit model, which accounted for the flexibility of enzymes. Under this new model, the active site, where the reactant binds, is viewed as dynamic. The amino acid side chains that make the active site are put into the positions necessary to perform catalysis, but are not necessarily rigid and can be shaped with the environment and the substrate. Moreover, according to this new model, the substrate can change shape slightly to enter the active site. In this way, the substrate and active site of the enzyme undergo slight conformation changes until full binding occurs and an affinity maximum is reached.
Figure 2 shows a picture of the induced fit model. We see here that the unbound enzyme active site does not perfectly complement the two substrates. However, upon substrate binding, we see that the global structure of the enzyme shifts, which causes an induced fit in which the active sites now complement the substrates.
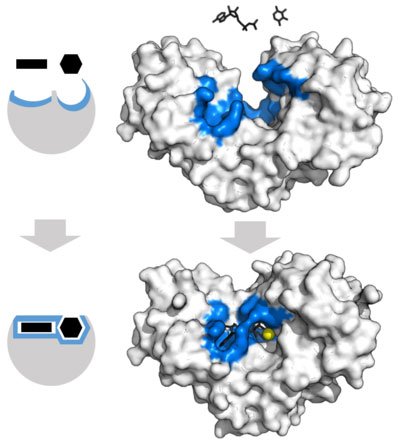
Figure 2: http://en.wikipedia.org/wiki/Enzyme#cite_note-Cooper_2000-34
These dynamic active sites not only complement the shape of the substrate, but its chemical nature as well. This allows enzymes to be highly selective between very similar molecules.
8
Take a simple example. Lactase, an enzyme needed to digest milk, cleaves lactose, a dissacharride, into glucose and galactose. Lactase can only cleave lactose, regardless of how similar other sugars may be to lactose. Sucrose happens to be one of those sugars. Also a disaccharide but made of fructose and glucose, sucrose is almost structurally identical to lactose. And yet, lactase cannot cleave sucrose; instead, the enzyme sucrase does this. The opposite is also true—sucrase cannot cleave lactose. The enzymes are specific for their substrates. While this may seem trivial, it explains how people become lactose intolerant. Insufficient lactase production results in an inability to process lactose resulting in lactose intolerance—other enzymes cannot perform lactase’s job.
Now take a more complicated example. With DNA replication, where a single mistake is propagated with each subsequent replication, specificity is necessary to sustain life.
DNA polymerase is one enzyme that proofreads the DNA. It checks that each specific DNA base is added correctly so that errors in the DNA occur less than 1 in 100 million base additions.
9
If this were not the case—if enzymes were not specific—mutations in the DNA would be rampant, mistakes would be propagated and death would result. Therefore, in the case of DNA production the ability of enzymes to be selective and accurate is crucial.
Enzyme Kinetics
Enzyme kinetics is the study of the rate of chemical reactions. In this field, the reaction rate is investigated in relation to numerous variables, such as substrate concentration, to determine any number of things, including the mechanism of the reaction, how activity of the enzyme is controlled, and how the enzyme is inhibited or promoted.
The presence of enzymes specifically alters the kinetics of a reaction. Enzymes do not alter the equilibrium position—the point at which the rates of the forward and backward reactions are equal. The enzyme, however, alters the rate of the reaction and introduces saturating behavior. When small amounts of substrate are introduced to enzyme, the reaction is free to proceed linearly with substrate concentration; as one increases the amount of substrate, the amount of product formed increases by a proportional amount. However, at high substrate concentrations, the amount of product formed does not increase by a proportional amount. Instead, the reaction rate slows and reaches a maximum. In this case, most of the enzyme active sites are occupied by, or saturated with, substrate, which limits the reaction to how quickly each enzyme can turnover the substrate. In biology, this turnover is particularly important. Enzymes are often naturally found in very low proportion to their reactants and as such, reactions can only proceed as quickly as reactants can be turned over.
Michaelis-Menten Kinetics
The Michaelis-Menten kinetic model is an important introduction to the realm of reaction rates and the concept of enzyme saturation. Initially, at lower substrate concentrations, the reaction rate increases linearly with substrate concentration. However, at some point, saturation occurs and the rate of reaction reaches a maximum even upon the addition of more substrate.
To illustrate the concept of saturation, the schematized model for an enzyme-catalyzed reaction is shown in Figure 3. It shows that there is an initial bimolecular reaction between enzyme E and substrate S, which forms the enzyme substrate complex ES. Here, we can see how saturation occurs. As enzyme reacts with substrate to form ES, fewer and fewer enzyme molecules are left to react with substrate. Eventually, all enzymes will be bound to ES and saturation will have occurred. Once saturation occurs, the rate of the reaction becomes dependent on the concentration of ES and is essentially a unimolecular reaction.
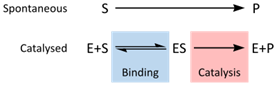
Figure 3: http://en.wikipedia.org/wiki/File:Enzyme_mechanism_2.svg
K
m
in the Michaelis-Menten Equation
The Michaelis-Menten equation is displayed below and describes how the initial reaction rate (V
0
) depends on the concentration of substrate ([S]):
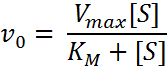
The K
m
in this equation is especially noteworthy. It is a useful index for affinity between the substrate and the enzyme. Specifically, K
m
serves as a dissociation constant such that larger K
m
values mean lower affinity between the substrate and enzyme and smaller K
m
values mean the higher affinity between the substrate and the enzyme. In other words, K
m
lets us know the affinity between the substrate and the enzyme.
10
To illustrate this, we will investigate reaction rates and their constants, indicated by K
1
, K
2
, and K
-1
. The enzyme-catalyzed reaction equation below shows the formation of product from substrate, facilitated by an enzyme, where E
f
corresponds to the free enzyme, ES corresponds to the enzyme-substrate complex, S corresponds to the substrate, and P corresponds to the product.
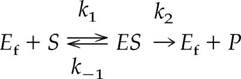
From the equation above, we see that the rate of dissociation corresponds to the two arrows pointing away from ES—K
-1
and K
2
. Additionally, we see that the rate of creation of ES corresponds to the one arrow pointing towards ES—K
1
. Dividing the rate of dissociation of ES by the rate of creation of ES, we get K
m
, as shown below.
K
m
= (K
-1
+K
2
)/K
1
In enzyme-catalyzed reactions, the rate-limiting step is generally the conversion of ES to free enzyme and product. Therefore, the arrow pointing from ES to E
f
and P—K
2
—represents a small value. In fact, the value of K
2
is so small in comparison to that of K
-1
that the equation can be simplified to:
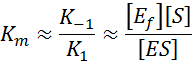
This shows that K
m
is an approximation of the dissociation constant. It indicates the affinity between the enzyme and the substrate such that high K
m
values indicate a low affinity and low K
m
values indicate a high affinity.
Lets explore K
m
further. The Michaelis-Menten equation shows that K
m
must have units of substrate concentration since it appears in the denominator (shown below). Therefore, there must exist a point at which K
m
equals the concentration of the substrate. To determine the point at which this occurs requires manipulation of the Michaelis-Menten equation. If you set [S] = K
m
in the below equation and simplify, you get the subsequent equation, which shows that, indeed, K
m
is the concentration of the substrate when V
0
is half of V
max
.
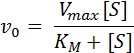
V
0
= V
max
/2
This finding—that K
m
is the concentration of the substrate when V
0
is half of V
max
—is particularly important because it allows us to determine K
m
by analyzing a saturation curve, like the one shown below.

Figure 4: https://en.wikipedia.org/wiki/Michaelis%E2%80%93Menten_kinetics#/media/File:Michaelis_Menten_curve_2.svg
Using this graph, we can see how K
m
is determined from the equation we derived. Knowing that K
m
is the concentration at which V
0
is half of V
max
, we can look at the graph above, find V
max
, take half of it, and then determine the substrate concentration corresponding to it. That concentration is K
m
or the dissociation constant.